cgRNASP-CN: a minimal coarse-grained representation-based statistical potential for RNA 3D structure evaluation
Ling Song,Shixiong Yu,Xunxun Wang,Ya-Lan Tanand Zhi-Jie Tan,*
1 Department of Physics and Key Laboratory of Artificial Micro&Nano-structures of Education,School of Physics and Technology, Wuhan University, Wuhan 430072, China
2 Research Center of Nonlinear Science, School of Mathematical and Physical Sciences, Wuhan Textile University, Wuhan 430073, China
Abstract Knowledge of RNA 3-dimensional (3D) structures is critical to understand the important biological functions of RNAs, and various models have been developed to predict RNA 3D structures in silico.However, there is still lack of a reliable and efficient statistical potential for RNA 3D structure evaluation.For this purpose, we developed a statistical potential based on a minimal coarse-grained representation and residue separation, where every nucleotide is represented by C4’ atom for backbone and N1 (or N9) atom for base.In analogy to the newly developed all-atom rsRNASP, cgRNASP-CN is composed of short-ranged and long-ranged potentials, and the short-ranged one was involved more subtly.The examination indicates that the performance of cgRNASP-CN is close to that of the all-atom rsRNASP and is superior to other top all-atom traditional statistical potentials and scoring functions trained from neural networks, for two realistic test datasets including the RNA-Puzzles dataset.Very importantly,cgRNASP-CN is about 100 times more efficient than existing all-atom statistical potentials/scoring functions including rsRNASP.cgRNASP-CN is available at website: https://github.com/Tan-group/cgRNASP-CN.
Keywords: RNA structure prediction, statistical potential, structure evaluation
1.Introduction
Noncoding RNAs have crucial biological functions such as regulating gene expression and catalyzing some biochemical reactions [1–4], and the functions of RNAs are generally correlated to their structures, especially three-dimensional(3D) structure [5, 6].Due to the high cost of experimental methods such as x-ray crystallography, NMR spectroscopy and cryo-electron microscopy, the high-resolution 3D structures of RNAs stored in protein database bank(PDB)are still very limited [7].Parallelly, some theoretical/computational models have been developed to predict the 3D structures of RNAs [8–14] either based on certain physical principles or based on existing structures in PDB database [7], and correspondingly the models can be roughly divided into physicsbased ones [15–22] and knowledge-based one [23–25].The physics-based models such as SimRNA [26, 27], IsRNA[28–30], iFold [31], NAST [32], HiRE-RNA [33], and our model of salt effect [34–40], are generally based on coarsegrained (CG) representations, specified CG force fields, and certain conformation sampling strategies.The knowledgebased models such as MC-fold/MC-sym pipeline, FARNA[25],Vfold3D[41–44],RNAComposer[45,46],and 3d RNA[47,48],are generally based on various fragment libraries and fragment-assembly strategies.Generally, an RNA 3D structure prediction model generally generates a large number of 3D structure candidates for a target RNA,and consequently,a reliable statistical potential/scoring function is required to identify a structure closest to the native one [49, 50].Furthermore, a reliable statistical potential can be involved in guiding RNA conformational sampling [26–30].
Knowledge-based statistical potentials have been shown to be rather effective and efficient in structure prediction and evaluation for proteins [51–57], protein-ligand complexes[58]and protein-protein complexes[59,60].There have been six kinds of reference states commonly used in building statistical potentials, i.e.average reference state [54], quasichemical approximate reference state [57], atomic-shuffle reference state [61], finite ideal-gas reference state [62],spherical non-interaction reference state [63] and randomwalk chain reference state [64].For RNA 3D structure evaluation,some statistical potentials have been developed based on different reference states [4, 65–67].Bernauer et al developed differentiable statistical potentials of KB at both all-atom and CG representations based on the quasi-chemical approximation reference state [65].Capriotti et al built allatom and CG statistical potentials of RASP based on the averaging reference state[66].Wang et al derived an all-atom distance and torsion-angle-dependent statistical potential of 3dRNAscore based on the average reference state [4].Zhang et al proposed an all-atom distance-dependent statistical potential of DFIRE based on the finite ideal-gas reference state [68].By building six statistical potentials based on the same training set and six existing reference states, we found that the finite ideal-gas and random-walk chain reference states are slightly better than other reference states in identifying native structures and ranking decoy structures [67].Recently, machine learning/deep learning approaches have been used in building scoring functions RNA 3D structure evaluation [69, 70].Compared with the top traditional statistical potentials, RNA3DCNN constructed by 3D convolutional neural network shows excellent performance in identifying native structures of RNA-Puzzle dataset[69],and the newly developed ARES [70], from deep neural network based on training data from FARFAR2 showed rather good performance for evaluating structures from FARFAR2 [71].Very recently, we have developed an all-atom residueseparation-based statistical potential of rsRNASP through distinguishing short-ranged and long-ranged potentials, and rsRNASP shows a visibly improved performance than existing statistical potentials and scoring functions from neural networks [72].
However, almost all existing physics-based models for RNA 3D structure prediction are based on different-level CG representations rather than the all-atom one to reduce conformational space, while the existing statistical potentials/scoring functions of high performance are all based on the allatom representation.Consequently, a reliable CG statistical potential is crucially important for a CG-based 3D structure prediction model rather than an all-atom-based one.Furthermore,a reliable CG statistical potential can also be applicable for all-atom structure evaluation at much higher efficiency than an all-atom one since much fewer CG atoms are involved.Therefore, reliable CG statistical potentials are still highly required,not only for CG structure evaluation but also for all-atom structure evaluation at high efficiency.
In this work, we developed a CG statistical potential of cgRNASP-CN for RNA 3D structure evaluation based on a minimal CG representation for nucleotides.Specifically, we used two real heavy atoms of C4’,and N1(for purines or N9 for pyrimidines) for describing a nucleotide, and C4’ atoms and N1 (or N9) atoms describe the backbone and bases for an RNA chain, respectively.The examinations for realistic datasets show that cgRNASP-CN has a good performance in structure evaluation for realistic test sets including the RNAPuzzles dataset.Furthermore, for the RNA-Puzzle dataset,the performance of cgRNASP-CN is very close to the newly developed all-atom rsRNASP and superior to other top allatom statistical potentials/scoring functions.Very importantly, cgRNASP-CN is (over) ~100 times more efficient than existing top all-atom statistical potentials/scoring functions.
2.Methods
A minimal CG representation
First, we surveyed the physics-based RNA 3D structure prediction models [26, 29, 38], and found that C4’ and N1 (for purines or N9 for pyrimidines) atoms were used very frequently for describing backbone and base for a nucleotide,respectively.For example, C4’ and N1 (or N9) atoms were used in SimRNA [26, 27], Vfold [41–44], Shapiro’s model[73], and our CG model with salt effect [34–40].Moreover,the CG representation of two atoms of C4’ and N1 (or N9)can be considered as a minimal one for describing backbone and base for a nucleotide, respectively; see figure 1.Thus,based on the minimal CG representation,we developed a new CG statistical potential, namely cgRNASP-CN, for RNA 3D structure evaluation.
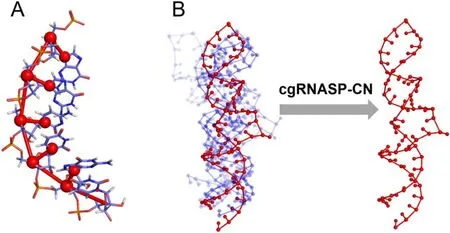
Figure 1.(A) Illustration of a minimal coarse-grained (CG) representation used in developing the statistical potential of cgRNASP-CN for RNA 3D structure evaluation,where 2 CG beads at C4’and N1/N9 heavy atoms describe backbone and base,respectively;(B)illustration for top-1 structure identified by cgRNASP-CN from the structure candidates.
A CG statistical potential based on a minimal CG representation and residue separation
RNA folding is generally hierarchical [74], and the interactions at different residue separations may play different roles in stabilizing RNA 3D structures [51].Similar to the newly developed all-atom rsRNASP [72], the total energy of an RNA conformation C of a given sequence is composed of short-ranged energy and long-ranged energy in the present cgRNASP-CN [75]:
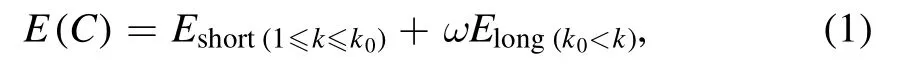
where k0is a residue separation threshold to distinguish shortand long-ranged interactions and ω is a weight to balance the two contributions.The long-ranged energy Elongin cgRNASP-CN can be given by [72, 75]

and
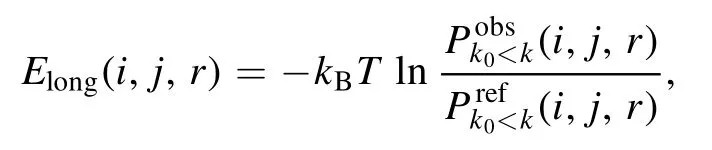
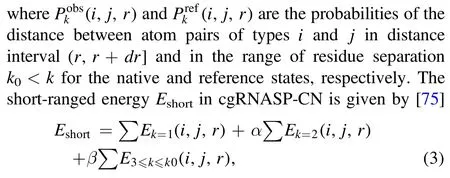
and
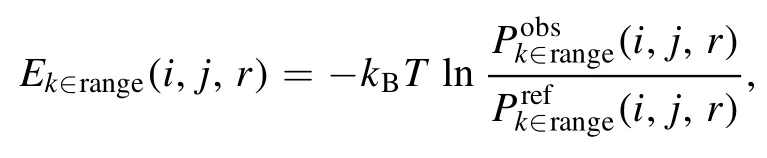
where k ∈ range stands for the residue separation k in the k
Training set and parameters
In cgRNASP-CN, we used the same non-redundant training native set that was recently used to derive allatom rsRNASP [72], and the dataset is available at https://github.com/Tan-group/rsRNASP [72].It should be noted that there are several RNAs in the training set with over 80% identity with the RNAs in the test set, and these RNAs were still reserved in the training set for keeping the complete structure spectrum.For these RNAs, we used the leave-oneout method according to previous works [67, 72, 75].To optimize the weights (α, β, ω) in equations (1) and (3) for short- and long-ranged interactions, we used a training decoy dataset previously built for deriving the all-atom rsRNASP,which is available at https://github.com/Tan-group/rsRNASP [72].According to all-atom rsRNASP [72],k0is taken as 4, and an RNA length N-dependent function f(N)was involved to normalize the N-dependent CG bead-pair number of the long-ranged interactions due to the large residue-separation range and the consequent N-dependent CG bead-pair number.Consequently,ωin equation(1)is equal toω=ω0/f(N).Based on the examinations on the training decoy dataset,α, β andω0were determined for cgRNASPCN, respectively.For the details of f(N) andα, β andω0,please see section S3 and figures S2,S3 in the supplementary material.
In cgRNASP-CN, the distance bin width is 0.3 ?[4, 67, 72, 75], and the cut-off distances of the statistical potential of 1, 2, 3–4 and 4<kin the k range are set to the values according to the distance distribution between CG beads in different residue-separation ranges [72]; please see section S2 and figure S1 in the supplementary materials for more detailed information.For the case that some atom-pairs are not observed within a certain bin width, the potentials were set to the highest potential value in the whole range for corresponding CG atom pair types, and when the distance of CG atomic pairs is less than 3.9 ? (mean van der Waals diameter for C4’ and N1 (or N9) atoms), the potentials were set to a high value of 50,wherekBTwas taken as the unit of potential energy.
Test datasets
In order to test the performance of cgRNASP-CN, we used two realistic test sets of the PM and Puzzles datasets instead of those from perturbation methods[72].The PM dataset was built by us previously through four RNA 3D structure prediction models with given native secondary structures, which is composed of decoy structures for 20 RNAs and is available at https://github.com/Tan-group/rsRNASP [72].The Puzzles dataset was generated from the CASP-like competition of RNA 3D structures predictions,and is composed of the decoy structures of 22 RNAs from various top research groups around the world [67].The Puzzles dataset is available at https://github.com/RNA-Puzzles/standardized_dataset, and the Puzzles dataset is of particular importance since it was generated from the blind CASP-like 3D RNA structure predictions from various top research groups with given sequences [67].
Measuring RNA structure similarity
To measure the structural difference between the two RNA 3D structures, we used both root-mean-square-deviation(RMSD) and deformation index (DI) metrics.The DI value between structures A and B is calculated as follows [76]:
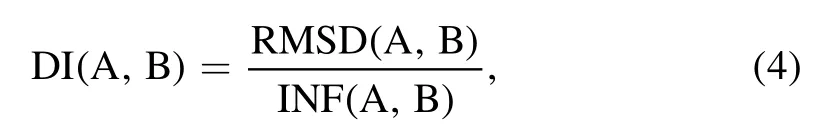
where RMSD (A , B) and INF (A , B) represent geometric and topological differences between structures A and B, respectively.INF describes interaction network fidelity and can be measured by Matthews correlation coefficient of base-pairing and base-stacking interactions[77].If structures A and B have very similar hydrogen bond interaction networks, the DI will be similar to the RMSD, otherwise the DI value will be relatively larger than the RMSD value.The tools for calculating DI and INF are available at https://github.com/RNAPuzzles/BasicAssessMetrics [78].
3.Results and discussion
In the following,we tested the performance of cgRNASP-CN against two realistic datasets PM and Puzzles,in a comparable way with existing top all-atom statistical potentials/scoring functions including rsRNASP[72],RNA3DCNN[69],ARES[70], DFIRE-RNA [68], 3dRNAscore [4], and RASP [66].First,we examined the overall performance of cgRNASP-CN against the two datasets, and afterwards focused on the performance of cgRNASP-CN against the RNA-Puzzles dataset.Finally, we examined the computation efficiency of cgRNASP-CN, compared with the existing top all-atom statistical potentials/scoring functions.
Evaluation metrics
To describe the performance of cgRNASP-CN, we used the following three metrics: (a) the number of identified native structures; (b) the DI values of the lowest-energy structure(including and excluding the native structure), and (c) the Pearson correlation coefficient (PCC) between energies and DIs of decoy structures.The PCC value is calculated as follows:
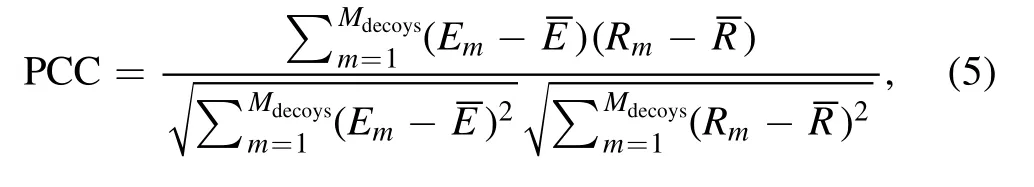
whereMdecoysis the total number of decoy structures for an RNA.EmandRmare the energy and DI of the mth decoy structure, respectively.andare the averaged energy and DI of all decoy structures,respectively.The PCC value ranges from 0 to 1, and when PCC is equal to 1, the statistical potential has a perfect performance.
Overall performance of cgRNASP-CN for PM and Puzzles datasets
In identifying native structures.As shown in figure 2(A) and table S2 in the supplementary material, cgRNASP-CN identifies 30 native structures from the decoys of 42 RNAs for the PM and Puzzles datasets,i.e.cgRNASP-CN identifies~71% native structures for the two realistic datasets.In contrast, rsRNASP, RNA3DCNN, ARES, DFIRE-RNA,3dRNAscore, and RASP identify 32, 27, 2, 20, 4, and 4 native structures from the decoys of 42 RNAs for the two datasets.This indicates that the performance of cgRNASPCN is slightly lower than the all-atom rsRNASP, while is higher than other all-atom statistical potentials/scoring functions in identifying native structures.
In identifying near-native structures.We also examined the performance of cgRNASP-CN in identifying near-native structures for the two realistic datasets involving native structures.As shown in figure 2(B) and table S2 in the supplementary material, the mean DI of lowest-energy structures from cgRNASP-CN is ~3.5 ? for the two realistic datasets with native structures.Such value becomes 3.9 ? for rsRNASP, 6.1 ? for RNA3DCNN,16.1 ? for ARES, 8.9 ? for DFIRE-RNA, 16.5 ? for 3dRNAscore, and 17.4 ? for RASP, respectively.Namely,the DI from cgRNASP-CN is slightly smaller than that from the all-atom rsRNASP, while is apparently smaller than those from other top all-atom statistical potentials/scoring functions including RNA3DCNN, ARES, DFIRE-RNA,3dRNAscore, and RASP.This indicates the overall better performance of cgRNASP-CN than other top all-atom statistical potentials and scoring functions in identifying near-native structures for the two realistic datasets involving native ones.
Furthermore, we examined the ability of cgRNASP-CN in identifying near-native structures for the two datasets without involving native ones, since a 3D prediction model generally cannot generate native structures.As shown in figure 2(C) and tables S2 in the supplementary material, the mean DI from cgRNASP-CN is 12.6 ?, a smaller value than those from other existing top statistical potentials and scoring functions,while such values are 12.8 ? for rsRNASP,15.6 ? for RNA3DCNN,17.1 ? for ARES,13.8 ? for DFIRE-RNA,18.3 ? for 3dRNAscore,and 19.4 ? for RASP.This suggests that cgRNASP-CN has very slightly better performance than rsRNASP and visibly better performance than RNA3DCNN,ARES,DFIRE-RNA,3dRNAscore,and RASP,in identifying near-native structures for the two realistic datasets without native ones.
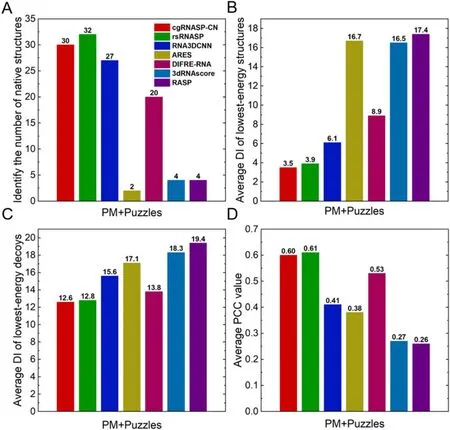
Figure 2.(A) Number of identified native structures, (B) average DI values of the lowest-energy structures (including native ones), (C)average DI values of the lowest-energy decoys(excluding native ones),and(D)average PCC values between DIs and energies by cgRNASPCN and other all-atom statistical potentials.Panels (A)–(D) are for the two realistic datasets (PM + Puzzles), and the PCC values were averaged over the mean values of respective test sets since decoys in a dataset were generated with the same method and have similar structure features.
In ranking decoy structures.A good statistical potential cannot only identify the near-native structures from decoys,but also rank the decoy structures according to their similarity to the native ones.We used the PCC between energies and DIs of decoys to assess the ability of cgRNASP-CN in ranking decoy structures of RNAs.As shown in figure 2(D)and table S2 in the supplementary material, the PCC value from cgRNASP-CN is ~0.60 for the two realistic datasets.Such value is very slightly smaller than that of rsRNASP(PCC ~ 0.61), while appears visibly larger than those of RNA3DCNN (PCC ~ 0.41), ARES (PCC ~ 0.38), DFIRERNA(PCC ~ 0.53),3dRNAscore (PCC ~ 0.27),and RASP(PCC ~ 0.26).Thus, cgRNASP-CN is very similar to rsRNASP while is visibly superior to other statistical potentials and scoring functions in ranking decoy structures.
Therefore, for the two realistic datasets, the present cgRNASP-CN is very similar to the all-atom rsRNASP and is visibly superior to other top all-atom statistical potentials/scoring functions for RNA 3D structure evaluation.It is encouraging that cgRNASP-CN appears very slightly better than rsRNASP in identifying near-native structures for the realistic PM and Puzzles datasets.
Performance of cgRNASP-CN for RNA-Puzzles dataset
The Puzzles dataset was generated from the CASP-like competition of RNA 3D structures predictions, and is composed of the decoy structures of 22 RNAs from various top research groups around the world.Due to the particular importance of the Puzzles dataset, in the following, we explicitly examined the performance of cgRNASP-CN against the Puzzles dataset.
In identifying native/near-native structures.As shown in figures 3(A)–(C) and table S3 in the supplementary material,cgRNASP-CN identifies 14 native structures from the decoys of 22 RNAs in the Puzzles dataset, and such number of identified native ones is slightly smaller than that of the all-atom rsRNASP (16 out of 22) while is larger than those of other all-atom statistical potentials/scoring functions including RNA3DCNN (13 out 22), ARES (2 out of 22),DFIRE-RNA (10 out of 22), 3dRNAscore (2 out of 22), and RASP (2 out of 22).Moreover, the DI values for the Puzzles dataset with and without native structures from cgRNASP-CN are 5.1 and 13.2 ?, which are similar to those from rsRNASP(4.6 and 14.4 ?) and appear smaller than those from RNA3DCNN (5.9 and 18.5 ?), ARES (18.1 and 18.8 ?),DFIRE-RNA (7.6 and 14.4 ?), 3dRNAscore (17.1 and 19.4 ?), and RASP (17.8 and 20.0 ?).This indicates that cgRNASP-CN is similar to the all-atom rsRNASP in identifying near-native structures and appears superior to other statistical potentials and scoring functions.Importantly,it is noted that the DI value of cgRNASP-CN for the Puzzles dataset without native structures is slightly smaller than that from rsRNASP, suggesting that cgRNASP-CN can identify structures closer to native ones than the all-atom rsRNASP since a native structure is generally absent for a blind structure prediction.
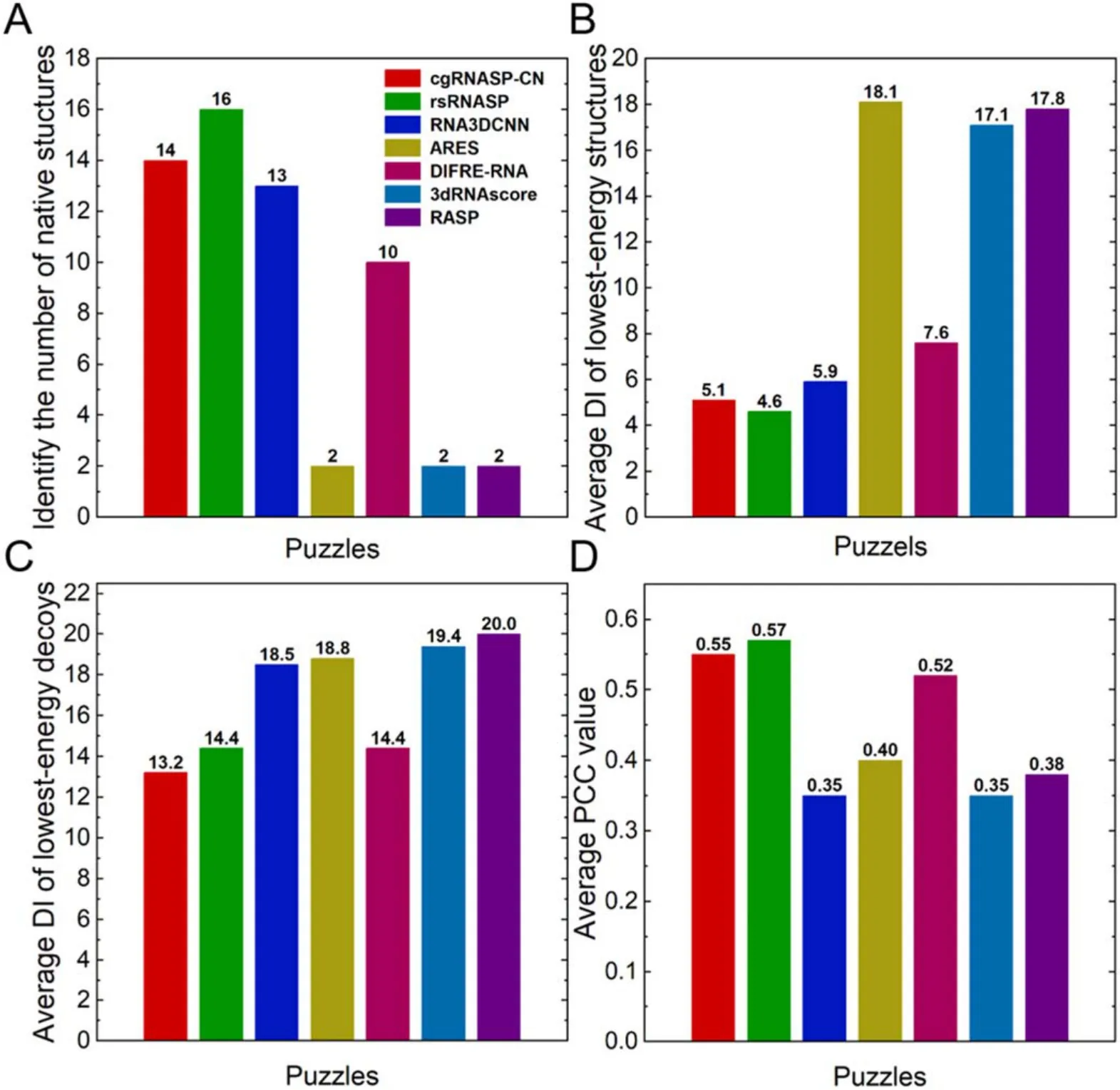
Figure 3.(A) Number of identified native structures, (B) average DI values of the lowest-energy structures (including native ones), (C)average DI values of the lowest-energy decoys (excluding native ones), and (D) average values of PCCs between DIs and energies by cgRNASP-CN and other all-atom statistical potentials for the Puzzles dataset.
In ranking decoy structures.The PCC values between energies and DIs of decoys of the Puzzles dataset are shown in figure 3(D) and tables S3 in the supplementary material.The PCC from cgRNASP-CN is 0.55, a slightly lower value than that from rsRNASP (0.57).However, the PCC from cgRNASP-CN is visibly higher than those from other top all-atom statistical potentials/scoring functions including RNA3DCNN (0.35), ARES (0.40), DFIRE-RNA(0.52), 3dRNAscore (0.35), and RASP (0.38).This suggests that cgRNASP-CN is close to rsRNASP and appears superior to other all-atom statistical potentials/scoring functions in ranking decoy structures for the Puzzles dataset.
Therefore, for the Puzzles dataset, the performance of cgRNASP-CN is overall similar to that of the all-atom rsRNASP and is better than other top all-atom statistical potentials/scoring functions.Notably, cgRNASP-CN can identify the structures closer to native ones when native structures are not involved in the Puzzles dataset, since a blind structure prediction generally does not involve a native structure.
Computation efficiency of cgRNASP-CN
As shown above, for the two realistic datasets of PM and Puzzles, the present cgRNASP-CN has a very similar performance with the newly developed all-atom rsRNASP and an overall better performance than other top all-atom statistical potentials/scoring functions.Since cgRNASP-CN is a statistical potential based on a minimal 2-bead CG representation, cgRNASP-CN can be employed not only for related CG structure evaluation, but also for all-atom structure evaluation at high efficiency.In the following, we quantitatively examined the computation efficiency of cgRNASP-CN for the RNAs in the Puzzles dataset, in a comparison with existing all-atom statistical potentials/scoring functions.
As shown in figure 4, for the RNAs in the Puzzles dataset,cgRNASP-CN is significantly more efficient than the all-atom statistical potential/scoring function of rsRNASP,RNA3DCNN,and DFIRE-RNA.Specifically,for the Puzzles dataset, the computation time of cgRNASP-CN is about 1/130 of that of rsRNASP, and the computation time of rsRNASP is comparable to that of DFIRE-RNA and is about 1/10 of that of RNA3DCNN.It is understandable since cgRNASP-CN involves a minimal 2-bead CG representation for a nucleotide and the computation time of a statistical potential generally is proportional to the square of atom number involved in the statistical potential.Therefore,cgRNASP-CN with good performance is significantly more efficient than existing all-atom statistical potentials/scoring functions,which would enable cgRNASP-CN to greatly save evaluation time for a given an ensemble of candidates or evaluate much more structure candidates within a given time.

Figure 4.Computation times of cgRNASP-CN and other top allatom statistical potentials/scoring functions for the Puzzles dataset containing decoys of 22 RNAs, relative to that of cgRNASP-CN.The PDB IDs of the 22 RNAs in the Puzzles dataset were shown as the X-axis label.
Conclusion
In this work, we developed the CG statistical potential of cgRNASP-CN based on a minimal CG representation for a nucleotide.The examinations against the realistic datasets show that compared with the newly developed all-atom rsRNASP, cgRNASP-CN has similar performance and even could identify nearer-native structures for the realistic datasets without involving native structures.Furthermore, cgRNASPCN is superior to other top existing all-atom statistical potentials/scoring functions for the realistic datasets.More importantly, cgRNASP-CN is significantly (over 100 times)more efficient than existing top all-atom statistical potentials/scoring functions including rsRNASP.Therefore, cgRNASPCN can be used not only for evaluating CG structure candidates with the corresponding CG atoms but also for evaluating all-atom structure candidates at very high efficiency.
However, the performance of cgRNASP-CN is still limited to a relatively good level.For example, for the realistic datasets,the percentage of identified native structures is~71% and the PCC value between DIs and energies of decoys is ~0.6,and such two values are still apparently lower than the ideal value of 1.Therefore,the present CG statistical potential of cgRNASP-CN is still required to be improved for a more reliable evaluation for RNA 3D structure candidates.First, due to the limited native RNA structures in the current PDB database [7], cgRNASP-CN can be continuously improved with the increase in the number of RNA structures deposited in the PDB database.Second, in addition to distance between CG atoms, some other geometric parameters such as torsion angle and orientation can be involved to develop a statistical potential to more completely capture the geometry of RNA 3D structures[4,24,79].Third,multi-body potentials can be explicitly involved in cgRNASP-CN,which will improve the description for correlated atom-atom distance distributions [24, 80].Nevertheless, the present statistical potential of cgRNASP-CN based on a minimal CG representation would be very beneficial for related CG-based 3D structure evaluation and for all-atom-based 3D structure evaluation at significantly high computation efficiency.
Acknowledgments
We are grateful to Profs Shi-Jie Chen(University of Missouri)and Jian Zhang(Nanjing University)for valuable discussions.The numerical calculations in this work were performed on the super computing system in the Super Computing Center of Wuhan University.
Data availability statement
All relevant data are within the paper and its supplementary material files.The potential of cgRNASP-CN is available at website https://github.com/Tan-group/cgRNASP-CN.
Author contributions
Z J T,Y L T and L S designed the research.L S,X W and S X Y performed the research.T Z J, Y L T, X W and L S analyzed the data.L S, Y L T, X W, and Z J T wrote the manuscript.
Funding
This work was supported by grants from the National Science Foundation of China (12075171, 11774272).
 Communications in Theoretical Physics2022年7期
Communications in Theoretical Physics2022年7期
- Communications in Theoretical Physics的其它文章
- Topological and dynamical phase transitions in the Su–Schrieffer–Heeger model with quasiperiodic and long-range hoppings
- Anisotropic and valley-resolved beamsplitter based on a tilted Dirac system
- Stable striped state in a rotating twodimensional spin–orbit coupled spin-1/2 Bose–Einstein condensate
- Density fluctuations of two-dimensional active-passive mixtures
- A new effective potential for deuteron
- The pseudoscalar meson and baryon octet interaction with strangeness S = -2 in the unitary coupled-channel approximation
